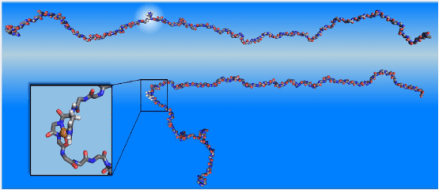
The protein alpha-synuclein in its normal state (above) and misfolded after the attachment of copper (below)
A team of physicists at North Carolina State University — led by my very good friend, Frisco Rose — has published the results of their study of the process that leads to Parkinson’s disease. Parkinson’s is a degenerative disorder that affects the nervous system, manifesting in tremors and difficulty controlling motion. Actors Katherine Hepburn and Michael J. Fox are well-known sufferers of the disease.
The work involved simulations using the most powerful supercomputer in the world, the Jaguar supercomputer at Oak Ridge National Laboratory, to understand the way in which a protein associated with Parkinson’s gets tangled. The protein, called alpha-synuclein, is normally long and straight, but it becomes tangled, or misfolded, in patients with Parkinson’s (see figure above).
Proteins are the basic building blocks of life. They are comprised of long chains of molecules called amino acids that regulate biochemical reactions in living things. The shape of a protein — the way in which it is folded — dictates its function. Amazingly, these biochemical machines assemble, or fold, themselves1.
Most of the time this self-assembly proceeds without error. However, when a protein misfolds, it becomes tangled and clumped together with other protein strands, and this is believed to cause a number of diseases, including Parkinson’s, Mad Cow, cystic fibrosis, and some forms of cancer. In order to devise treatments for these diseases, it’s important to understand how certain proteins misfold.
Study of protein folding may sound like a job for biologists, but it has been an increasingly popular topic of study in physics, because the different ways in which a protein can fold are determined by equations involving forces and energy. For a typical protein, these calculations would normally require hundreds of thousands of computing hours, far more than is feasible. To get around this problem, Rose’s team devised a new tactic: focus the simulations only on the part of the protein where the tangling occurs. By reducing the region of study, they were able to successfully carry out simulations, and discovered that certain metals, such as copper, affect the folding by binding to the protein in a way that accelerates tangling.
“We knew that the copper was interacting with a certain section of the protein, but we didn’t have a model for what was happening on the atomic level,” says Frisco Rose, Ph.D. candidate in physics and lead author of the paper describing the research. “Think of a huge swing set, with kids all swinging and holding hands—that’s the protein. Copper is a kid who wants a swing. There are a number of ways that copper could grab a swing, or bind to the protein, and each of those ways would affect all of the other kids on the swing set differently. We wanted to find the specific binding process that leads to misfolding.”
The tactic allowed them to identify the most likely way in which copper binding to the protein leads to misfolding. This is a significant step toward finding a treatment for Parkinson’s.
Other researchers studying protein folding are getting around the computing problem using a different tactic: using thousands of volunteered home computers — your computers — to perform the calculations. If you’d like to get involved by donating some of your home computer’s run time to help these scientists do their work, check out Standford University’s Folding@home project.
[1] Proteins synthesize and assemble themselves amazingly quickly, on the order of minutes. Given the truly astronomical number of possible configurations of molecules, it’s astounding that they are able to assemble so quickly into the proper configurations, i.e. the ones necessary for the existence of life as we know it. In other words, it’s not a process dictated by chance. It’s as though proteins “know” their correct configuration, and scientists are trying to understand how. See Chapter 7 of Gerald Schroeder’s The Science of God for a possible explanation.
Recommended reading:
